A Pilot Study to Investigate the potential for developing syndromic surveillance system based on meat inspection records in Western Kenya
A Pilot Study to Investigate the potential for developing syndromic surveillance system based on meat inspection records in Western Kenya

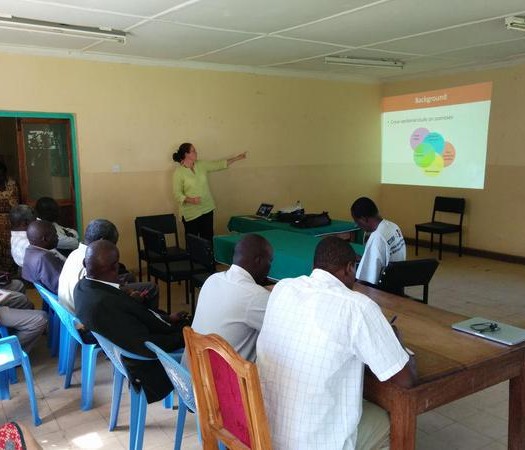
Training of meat inspectors on how to use hand held device for syndromic surveillance
Article by Joseph Ogola, ZooLinK Consultant
During our field visits in preparation for the ZooLink research project, we selected Kimilili and Webuye slaughterhouses in Bungoma County to participate in the syndromic surveillance pilot study. The two facilities within the study area were identified based on infrastructure and the willingness of the two meat inspectors to participate in the project. The rationale of this pilot project is to assess the feasibility of using slaughterhouse data to enhance the coverage and efficiency of the surveillance system in the study area alongside the routine laboratory based surveillance system. We developed a data collection form from the monthly reports from meat inspection records which
The rationale of this pilot project is to assess the feasibility of using slaughterhouse data to enhance the coverage and efficiency of the surveillance system in the study area alongside the routine laboratory based surveillance system. We developed a data collection form from the monthly reports from meat inspection records which
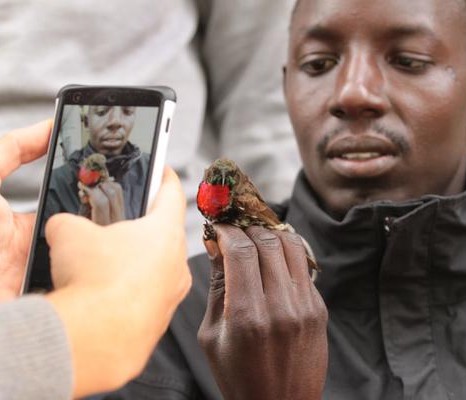
We developed a data collection form from the monthly reports from meat inspection records which were loaded onto a handheld device. The form captures information related to the carcass inspection together with animal location and movement data. The two meat inspectors after a short training session were then provided with two mobile phones to use daily to record data
(including any relevant photos) of animals slaughtered over a 6 month period. The data collected are sent directly to our data management platform.
We look forward to share the outcomes of this study in subsequent editions of the newsletter!








































You must be logged in to post a comment.